Martha M. Howe, PhD
Hariett S. Van Vleet Professor of Virology
858 Madison Ave.
Room 701 Molecular Science Building
Memphis, TN 38163
Email: mhowe@uthsc.edu
Phone: 901.448.8215
Fax: 901.448.7360
Research Interests
Turning genes on and off is a major regulatory mechanism used throughout the entire spectrum of biological organisms. Our laboratory is studying how genes get turned on and off in a model system based on the Mu phage, a virus that infects several kinds of bacteria, including Escherichia coli. Mu phage, like many viruses, turns its genes on in a defined order during development; first the early genes are expressed, then the middle genes, and then the late genes. Early gene expression occurs due to recognition of the phage early promoter by the host RNA polymerase (RNAP). The middle and late promoters cannot be recognized directly by this polymerase; they require an accessory DNA binding protein called an activator to bind to the promoter and help RNA polymerase recognize, bind, and activate gene expression from the promoter. Current efforts in the Howe laboratory are focused on dissecting the mechanism of transcription activation by the founding members of the novel Mor/C family of Class II activator proteins. Mor and C are the closely-related activators required for middle and late transcription, respectively, during lytic development of phage Mu. Our earlier work involved extensive mutagenesis of one middle (Pm) and one late (Plys) promoter and determination of the effects of those mutations on Mor or C binding.
These experiments showed that Mor and C bind, probably as dimers, to dyad-symmetry elements centered at -43.5 in the promoter. They also revealed that bases just upstream and downstream of the activator binding site influence promoter activity without affecting binding of the activator. In vitro transcription experiments with reconstituted mutant RNAPs showed that both the alpha-CTD and sigma-CTD are required for Mor-dependent activation of the middle promoter, leading to our working hypothesis for activation shown in Figure 1. Random and alanine-scanning mutagenesis of the C protein demonstrated that a predicted helix-turn-helix motif near the C terminus is required for DNA binding. Mutagenesis of the middle promoter and a model late promoter revealed that both contain an Up-like element that binds the alpha-CTD of RNA polymerase. From “order of addition” experiments, it appears that both C and Mor are able to recruit RNAP to the promoter, using alpha-CTD/UP-element, sigma/-10 element, and RNAP-activator interactions.
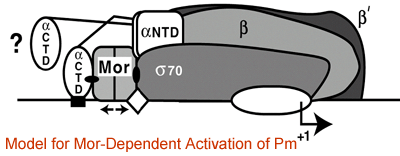
Our recent determination of the Mor structure by X-ray crystallography (in collaboration with Dr. Hee-Won Park in the Department of Structural Biology at St. Jude Children’s Research Hospital) revealed a dimer composed of two Mor monomers which each contain two domains, an N-terminal dimerization domain and a C-terminal DNA-binding domain; these domains are connected by a linker containing a beta strand. The N-terminal dimerization domains intertwine to form a four-helix bundle, generating a hydrophobic core that is further stabilized by antiparallel interactions between the two beta strands (Fig. 2).
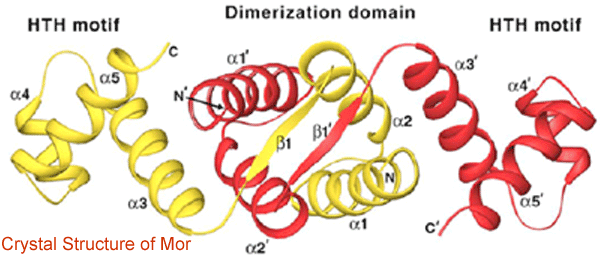
The C-terminal domain has a classical helix-turn-helix DNA-binding motif that is located at opposite ends of the elongated dimer. Since the distance between the two helix-turn-helix motifs is too great to allow binding to two adjacent major grooves of the 16-bp Mor-binding site, it is likely that conformational changes in both the protein and DNA will be required for Mor to interact with the DNA (Fig. 3).
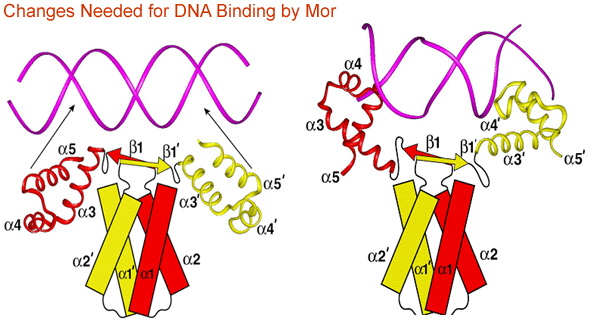
Experiments to test the key predictions from the structure are currently underway. Future experiments will include identification of specific amino acids involved in activator/RNAP interactions, determination of the role of promoter sequences flanking the activator binding site, and characterization of the conformational changes that occur in both the activator protein and promoter DNA.
"This material is based upon work supported by the National Science Foundation under
Grant No. 0418108.
Any opinions, findings, and conclusions or recommendations expressed in this material
are those of the
author(s) and do not necessarily reflect the views of the National Science Foundation."
PubMed search for Dr. Martha Howe
Education
- Bryn Mawr College, Bryn Mawr PA AB 1966 Biology
- Massachusetts Institute of Technology PhD 1972 Biology-Microbiology
- Centro Degli Acidi Nucleici, Rome, Italy postdoc 1972 Mu molecular biology
- Cold Spring Harbor Laboratory, NY postdoc 1973-4 Mu molecular biology
